Enzyme Indicated in Progression of Acute Myeloid Leukemia
Molecular mechanisms behind acute myeloid leukemia may offer a new target for drug development.
Molecular mechanisms behind acute myeloid leukemia may offer a new target for drug development.
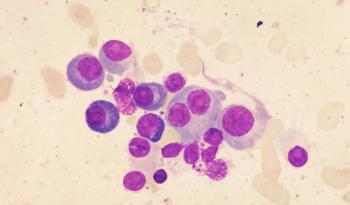
Acute myeloid leukemia is in particular need of new treatment options for patients. The malignancy begins in the bone marrow and can spread rapidly to the bloodstream, depriving the body of the essential blood cells that carry oxygen and fight infections.
New research from a team led by Rockefeller University scientists found a potential genetic weakness of the disease, offering insights into the molecular mechanisms behind acute myeloid leukemia and suggesting a new target for drug development.
Prior research indicates a variety of mutations are associated with the disease, including a DNA rearrangement found in about 15% of patients. This abnormal DNA-binding protein takes on new responsibilities, dramatically changing the way a set of genes that are turned on in a cell to promote the cancer.
But the reason why the mutation affects these changes is still a mystery to scientists.
A research team began to search for proteins that interact with a mutant protein known as AE, produced by a DNA rearrangement. They identified an enzyme that removes chemical tags, known as methyl groups from histones, which are proteins contained in chromosomes.
The enzyme is called JMJD1C. The tags act as repressive marks, suggesting that genes in the associated region should be turned off. Scientists wanted to explore the relationship between JMJD1C and AE, so they examined the broader effects of removing JMJD1C.
“We found that numerous genes were down-regulated upon loss of JMJD1C, and the set overlaps significantly with the genes that are normally activated by AE,” explained first author Mo Chen, a postdoc in Roeder’s lab.
As it turns out, the loss of gene expression has a dramatic consequence for the disease. The team discovered that acute myeloid leukemia cells are addicted to the presence of JMJD1C, and without it they cannot survive.
“In fact, these cell were very sensitive to depletion of JMJD1C,” Chen said. “We see an increase in apoptosis, a sort of cellular suicide.”
It was confirmed by scientists in the study that JMJD1C interacts with AE, and they demonstrated that the enzyme is required for AE to exert its cancer-promoting effects. The enzyme also plays an even broader role in acute myeloid leukemia beyond its interaction with AE, according to Chen.
“We were very surprised to find that JMJD1C is required for the proliferation of other acute myeloid leukemia cell lines, which do not have AE, so we looked for other proteins that might be responsible for JMJD1C addiction,” Chen said. The team found at least two other proteins that can recruit JMJD1C to target genes in diseased cells that lack AE, fueling leukemia growth.
The results indicate that JMJD1C may play a general role in promoting growth in myeloid leukemias.
“We are excited because this type of general phenomena is an ideal target for drug development,” Roeder said. “Our work will facilitate the development of selective inhibitors against JMJD1C, which is a highly promising therapeutic target for multiple types of leukemia.”