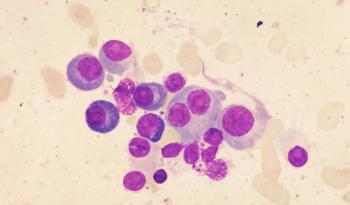
Mass Spectrometry Experiments Help Scientists See Effects of Drugs on Cancer Cells
Research offers enhanced method to assess effects of cancer drugs on normal and healthy cells.
Research offers enhanced method to assess effects of cancer drugs on normal and healthy cells.
Using discovery mass spectrometry (MS) experiments, researchers at EMBL’s European Bioinformatics Institute (EMBL-EBI) were able to develop a new method for studying the targets and effects of cancer drugs on normal and healthy cells.
It is difficult to fully comprehend the biological signaling pathways that regulate metabolism and gene expression because so many things are happening simultaneously. But understanding this process is crucial to knowing how a drug will affect healthy and cancer cells.
Protein kinases play an important role in these pathways as they turn certain proteins on and off during a process called phosphorylation. Because such pathways are often deregulated in cancer, kinase inhibitors are used as treatments.
Researchers in the study used data from ‘discovery mass spectrometry (MS) phospho-proteomics’ to create models of signaling pathways and networks. The models could be used to computationally test the effects of a drug.
“MS is a very powerful tool because it lets you look at activities like phosphorylation in a huge number of proteins at once,” said Julio Saez-Rodriguez, visiting group leader at EMBL-EBI. “But the data is typically very noisy, and random in terms of which of the tens of thousands of proteins in the cells it measures. It’s a little like trying to make a London tube map based on random CCTV photos.”
Until now, researchers have used methods using MS phospho-proteomics, leading them to find groups of kinases that are likely to be active in a sample. However, the new method takes it a step further by allowing scientists to ask more detailed questions about the exact effects the drug will have on proteins and pathways.
“Our method takes the noisy data from MS experiment, which is this large network with many interconnected cascades of kinase activities, filters the noise, and integrates the data. This is done entirely in the context of what we know about kinases and their substrates, so you can see how things are connected,” explained Camille Terfve, an EMBL International PhD student in Saez-Rodriguez’s lab. “Then you can compare what happens with the signal, for example if you use one kind of inhibitor or another. So, it can show what the drug is really doing to the system, beyond the direction you initially believed it would take.”
A collaboration with the Cutillas lab at the Barts Cancer Institute, Queen Mary University of London, allowed researchers to use data from experiments using kinase inhibitors on breast cancer cells to demonstrate the method.
“There is a lot of knowledge out there about protein kinases and how they influence phosphorylation, and we’ve put this together with the enormous potential of MS to create and test logic models that provide a clear path for research,” Saez-Rodriguez said. “MS produces so much data that’s very difficult to sift through — now, we can start to see the forest for the trees, and define which information is really important.”